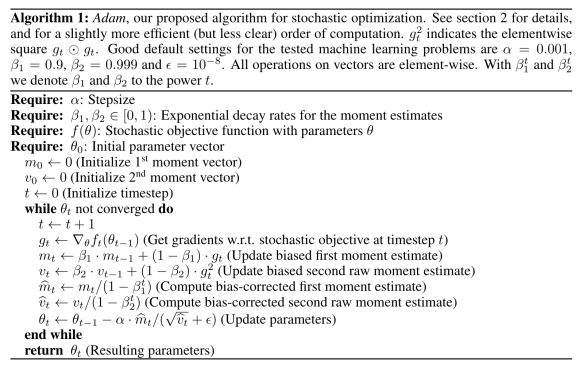
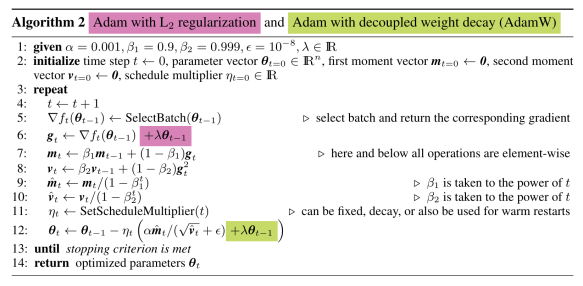
I recently learned from Allen Downey’s blog that Our World in Data is providing API access to their data. Our World in Data hosts datasets across several important topics, from population and demographic change, poverty and economic development, to human rights and democracy. From Nov 2024, Our World in Data “[offers] direct URLs to access data in CSV format and comprehensive metadata in JSON format” (this is what they call the Public Chart API).
See this link for full documentation on the chart data API. Allen Downey’s blog post shows how to use this API in Python; in this post I’ll show the corresponding code in R.
Downloading metadata
The chart data API page tells us what APIs are available to us. The starting point is a base URL which we denote by <base_url>. This is what’s available to us:
<base_url> – The page on the website where you can see the chart.<base_url>.csv – The data for this chart, usually what we want to download.<base_url>.metadata.json – The metadata for this chart, e.g. chart title, the units, how to cite the data sources.<base_url>.zip – The dataset, metadata, and a readme as zip file.
Let’s start by looking at the metadata for average monthly surface temperature:
library(tidyverse)
library(httr)
library(jsonlite)
# define URL and query parameters
url <- "https://ourworldindata.org/grapher/average-monthly-surface-temperature.metadata.json"
query_params <- list(
v = "1",
csvType = "full",
useColumnShortNames = "true"
)
# get metadata
headers <- add_headers(`User-Agent` = "Our World In Data data fetch/1.0")
response <- GET(url, query = query_params, headers)
metadata <- fromJSON(content(response, as = "text", encoding = "UTF-8"))
The returned metadata object is a list with a few keys:
# view the metadata keys
names(metadata)
#> [1] "chart" "columns" "dateDownloaded" "activeFilters"
The chart element gives us high-level information of the chart:
glimpse(metadata$chart)
#> List of 5
#> $ title : chr "Average monthly surface temperature"
#> $ subtitle : chr "The temperature of the air measured 2 meters above the ground, encompassing land, sea, and in-land water surfaces."
#> $ citation : chr "Contains modified Copernicus Climate Change Service information (2025)"
#> $ originalChartUrl: chr "https://ourworldindata.org/grapher/average-monthly-surface-temperature?v=1&csvType=full&useColumnShortNames=true"
#> $ selection : chr "World"
There’s more information on the dataset in metadata$columns$temperature_2m
names(metadata$columns$temperature_2m)
#> [1] "titleShort" "titleLong" "descriptionShort" "descriptionProcessing" "shortUnit" "unit"
#> [7] "timespan" "type" "owidVariableId" "shortName" "lastUpdated" "citationShort"
#> [13] "citationLong" "fullMetadata"
(Why temperature_2m? Looking at metadata$columns$temperature_2m$shortName, it seems like temperature_2m is the short name for the dataset.)
Downloading the dataset
Here is code downloading the average monthly surface temperature for just the USA:
# define the URL and query parameters
url <- "https://ourworldindata.org/grapher/average-monthly-surface-temperature.csv"
query_params <- list(
v = "1",
csvType = "filtered",
useColumnShortNames = "true",
tab = "chart",
country = "USA"
)
# get data
headers <- add_headers(`User-Agent` = "Our World In Data data fetch/1.0")
response <- GET(url, query = query_params, headers)
df <- read_csv(content(response, as = "text", encoding = "UTF-8"))
head(df)
#> # A tibble: 6 × 6
#> Entity Code year Day temperature_2m...5 temperature_2m...6
#> <chr> <chr> <dbl> <date> <dbl> <dbl>
#> 1 United States USA 1941 1941-12-15 -1.88 8.02
#> 2 United States USA 1942 1942-01-15 -4.78 7.85
#> 3 United States USA 1942 1942-02-15 -3.87 7.85
#> 4 United States USA 1942 1942-03-15 0.0978 7.85
#> 5 United States USA 1942 1942-04-15 7.54 7.85
#> 6 United States USA 1942 1942-05-15 12.1 7.85
Allen Downey notes that the last column is undocumented, and based on context is probably the average annual temperature.
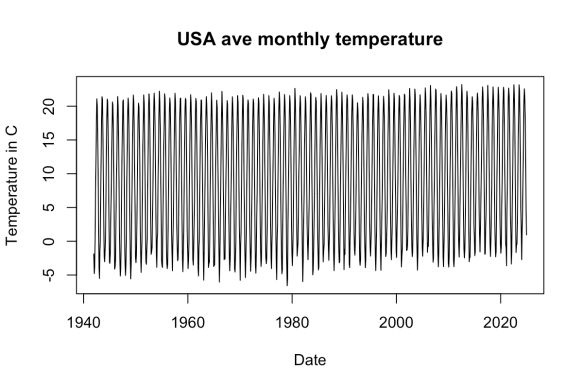
Here’s a simple plot of the data:
plot(df$Day, df$temperature_2m...5, type = "l",
main = "USA ave monthly temperature",
xlab = "Date", ylab = "Temperature in C")
If you wanted the entire dataset, just use csvType = "full" in the request parameters, like so:
# define the URL and query parameters
url <- "https://ourworldindata.org/grapher/average-monthly-surface-temperature.csv"
query_params <- list(
v = "1",
csvType = "full",
useColumnShortNames = "true",
tab = "chart"
)
# same code for the http request works, see above
How do you know which filters you can apply? There is no extensive documentation of what filters you can apply, but as you click on the base URL to change the chart, the URL changes. You can then use this to infer the query parameters to add.
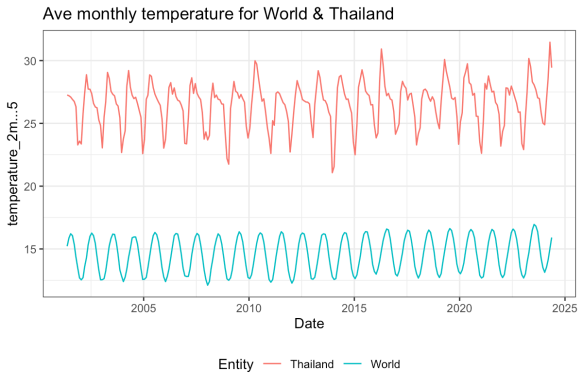
For example, I clicked around on the chart to get this URL: https://ourworldindata.org/grapher/average-monthly-surface-temperature?tab=chart&time=2001-05-15..2024-05-15&country=OWID_WRL~THA. The code below shows how you can download the data and reproduce the chart:
# define the URL and query parameters
url <- "https://ourworldindata.org/grapher/average-monthly-surface-temperature.csv"
query_params <- list(
v = "1",
csvType = "filtered",
useColumnShortNames = "true",
tab = "chart",
time = "2001-05-15..2024-05-15",
country = "OWID_WRL~THA"
)
# get data
headers <- add_headers(`User-Agent` = "Our World In Data data fetch/1.0")
response <- GET(url, query = query_params, headers)
df <- read_csv(content(response, as = "text", encoding = "UTF-8"))
ggplot(df) +
geom_line(aes(x = Day, y = temperature_2m...5, col = Entity)) +
labs(title = "Ave monthly temperature for World & Thailand",
x = "Date", y = "Temperature in C") +
theme_bw() +
theme(legend.position = "bottom")
is a discrete random variable taking values in
, then the probability-generating function (PGF) of
is defined as
is the probability mass function of
.
, and
,





