Conversation
|
For future reference, here are the 2 notebooks used when adding the new training reactions and refitting the rate tree |
16d59ab to
d5ae53b
Compare
|
Thank you for the update and for sharing the notebook for reviewing, @kspieks. It looks good. The previous root entry's parameters look so floppy (to me, A=4e-45, n = 16.9, E=-859 J/mol), and I am happy your recent update makes it more reasonable. In Kevin's current addition, it includes the reaction above. This definitely matches the template, but to me, it sounds more reasonable to be categorized to the [1,3]-sigmatropic rearrangement other than keto-enol according to my chemistry sense. While my point is maybe the scope of ketoenol is too broad or the name ketoenol is too specific. Moreover, I briefly checked the family list: we have [1,5]-H-shift as |
|
So in general I'm a fan of us merging families with close templates whenever possible. RMG just has so many families already to know and maintain. Additionally I think two families with similar templates will take almost twice as long to react as one family. Rate estimation is theoretically improved as well by merging families, but I think in practice it probably doesn't make that much of a difference in common cases we would do so. Although if there's some particular nuances in defining forbidden groups for these families I could see multiple families being justified. My opinion is that unless there's a particular nuance in the template or forbidden groups we should just keep it general and rename the family to something more appropriate, probably note that this family includes ketoenol reactions. Long term this reduces the amount of code we have to worry about. It may add a little bit of confusion, but there's already a lot of confusion about how named reaction types in literature map to RMG reaction types, we've really made that something people need to learn anyway if they want to touch the families in the database. |
|
@xiaoruiDong @mjohnson541 Thank you both for the discussion. To summarize ideas from our subgroups today, I will rebase this PR so these "traditional keto-enol" H transfer reactions can be merged in. Then I will create another PR that merges in other reactions that fit our template i.e. involve heavy atom transfers as R swaps with O. Then I will create a final PR that broadens this template by changing O to R i.e. We can name this new family 1,3-sigmatropic rearrangement and leave a note that it also includes keto-enol reactions. I will also add a few more training reactions to this last PR that match the new template |
d5ae53b to
1756fa9
Compare
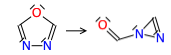
This PR adds 3 training reactions to the keto-enol family. All calculations were done at
CCSD(T)-F12/cc-pVDZ-F12//ωB97X-D3/def2-TZVP. The rate tree was then refit using Matt's notebook (#490).The long description includes the indices that label these species in the dataset paper I'm working on. These indices can be used to find the raw log files from the QM calculations.