-
Notifications
You must be signed in to change notification settings - Fork 252
Closed
Description
Topic
General area which your question is related to.
- Installation of RMG
- Running an RMG job
- Using RMG API
- Arkane (formerly CanTherm)
- Dependencies
- An error message
input file
#Data sources
database(
thermoLibraries =['BurkeH2O2','FFCM1(-)','thermo_DFT_CCSDTF12_BAC','CBS_QB3_1dHR','DFT_QCI_thermo','primaryThermoLibrary'], # 'FFCM1(-)','primaryThermoLibrary', 'BurkeH2O2','DFT_QCI_thermo','CBS_QB3_1dHR'
reactionLibraries = [('2005_Senosiain_OH_C2H2',False),('Glarborg/C3', False)], #
seedMechanisms = ['BurkeH2O2inN2','FFCM1(-)',], #
kineticsDepositories = ['training'],
kineticsFamilies = ['default'],
kineticsEstimator = 'rate rules',
)
#Constraints on generated species
generatedSpeciesConstraints(
maximumRadicalElectrons = 2,
allowed=['input species','seed mechanisms','reaction libraries'],
maximumCarbonAtoms=9,
maximumOxygenAtoms=4,
)
#List of species
species(
label='C7H16',
reactive=True,
structure=SMILES("CCCCCCC"),
)
species(
label='O2',
reactive=True,
structure=adjacencyList(
"""
1 O u1 p2 {2,S}
2 O u1 p2 {1,S}
"""),
)
species(
label='N2',
reactive=False,
structure=SMILES("N#N")
)
#Reaction systems
simpleReactor(
temperature=[(600,'K'),(1500,'K')],
pressure=[(10.0,'bar'),(40.0,'bar')],
nSims=8,
initialMoleFractions={
#"C7H10": 1,
"O2": 0.10885, # phi=1 means 9.5 O2 per C7H10
"N2": 0.881684808, # 8.1 times as much N2 as O2
"C7H16":0.00946, #
},
terminationTime = (1.0, 's'),
terminationRateRatio = 0.01,
terminationConversion={
'C7H16': 0.99,
},
)
simulator(
atol=1e-16,
rtol=1e-8,
)
model(
toleranceKeepInEdge=0, # No pruning to start
toleranceMoveToCore=0.4,
toleranceInterruptSimulation=1,
maxNumObjsPerIter=3,
terminateAtMaxObjects=True,
maxNumSpecies=200, # first stage is until core reaches 100 species
filterReactions=True, # should speed up model generation
)
model(
toleranceMoveToCore=0.25,
toleranceInterruptSimulation=1e8,
toleranceKeepInEdge=0.01, # Pruning enabled for stage 2
maximumEdgeSpecies=200000,
minCoreSizeForPrune=100,
minSpeciesExistIterationsForPrune=2,
filterReactions=True,
)
options(
units='si',
# saveRestartPeriod=(1,'hour'),
generateOutputHTML=False,
generatePlots=True,
saveSimulationProfiles=True,
saveEdgeSpecies=False,
)
Question
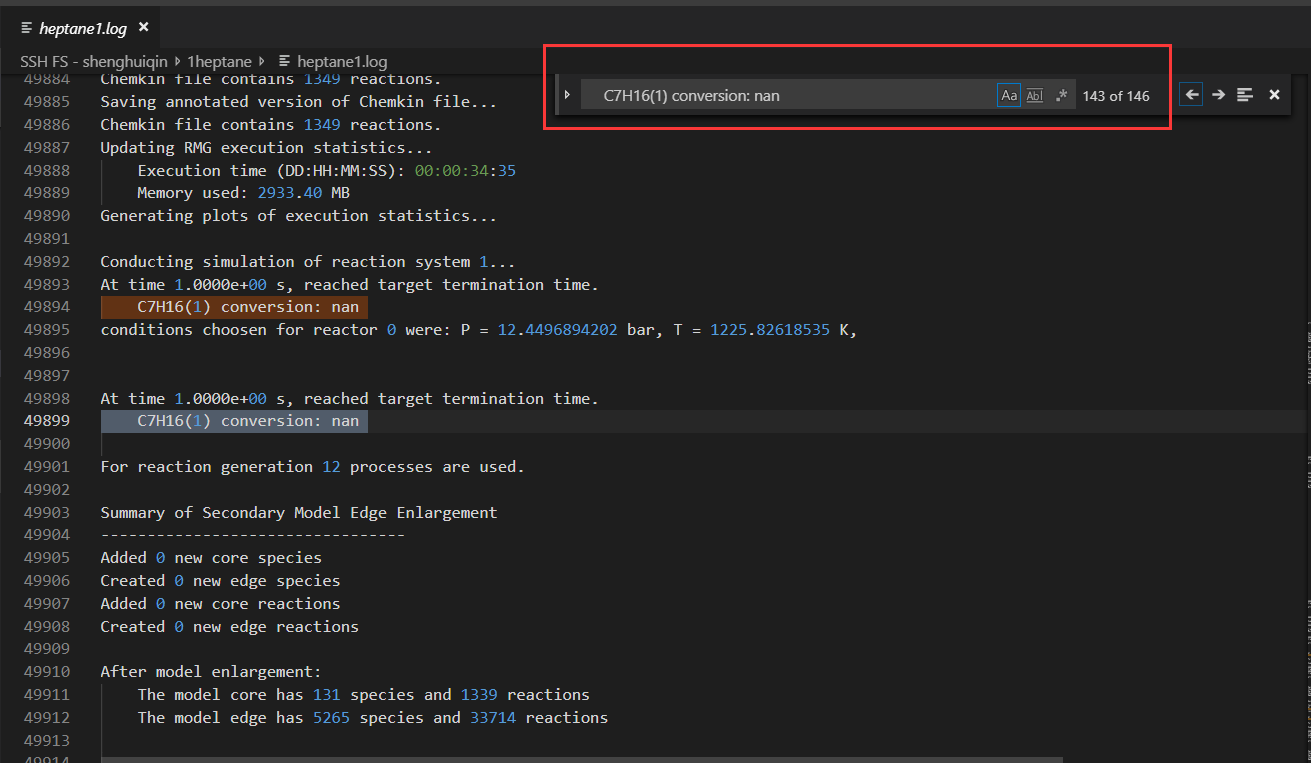
what happened, I get so many nan?
Usually, I won't get any nan with the similar input file, I want to know is this an error? something wrong with my input? or it's normal and I can ignore it,
Installation Information
Describe your installation method and system information if applicable.
- OS (include version if known): ubuntu
- Installation method: source install using conda env
- RMG version information:
- RMG-Py: july 31
- RMG-database: july 26
Reactions are currently unavailable
Metadata
Metadata
Assignees
Labels
No labels