Conversation
|
The process is written from a use-case perspective based on an example from @clausmichele. My question is whether this can be implemented by our back-ends, so for Platform, this is especially aimed at @jdries and @lforesta. |
|
Hi @m-mohr, I'm trying to implement fit_curve, but I don't understand how to write the fitting function. I will use dimension_labels to get the timestamps, is it ok in your opinion? |
|
@m-mohr could you please have a look here? I can't proceed with the implementation otherwise |
|
What is PS: I was on vacation last week and I'm still catching up. |
|
parameters is an array of coefficients, for this specific case is [a0,a1,a2], because the function I need to fit is: |
# Conflicts: # CHANGELOG.md
|
I've pushed an update to the process specification which adds a "parameters" parameter, which we (may?) need to define the number of parameters. Here's how a process graph could look like for the formula you gave: Please ensure that dimension labels for the temporal dimension are actually numerical and not just ISO strings! PS: This is purely theoretical and I've never actually tried or implemented it, so happy to take any suggestions that may make it easier or better etc. |
|
Thanks. I was actually thinking that the fit_curve process takes care of translating the temporal labels/strings (or whatever format they are) into the format required by the process (numerical). |
|
If that's a common use case we could think about this special case and support it, yes. |
|
Hmm, I'm just thinking whether it would be easier if this process returns an actual process graph instead of just the parameters? |
|
With the computed parameters [a0,a1,a2] and new time steps coming from another "testing" datacube (load_collection with a subsequent time extent) I need to predict the values the fitted function would take with the new time steps. This would basically return the a datacube with the same temporal extent as the "test" one, but different values: no more real data but predicted values. |
|
Results from a call with @clausmichele:
|
|
@clausmichele I made an attempt to cover what we discussed in a single process, which returns 2(!) data cubes. I have not had the time to fine-tune the descriptions yet, but would you think it would work this way for you? |
|
How would I have to use the result? How do I select the "parameters" datacube to predict values for a different datacube with different timesteps (same x,y,bands)? |
You get the data cube with
As above, but use 1 instead of 0.
Not possible. That use case didn't came up yet or I did not understand it.
I'm not sure what you want to have. You input a data cube (e.g. S2 with dimensions x,y,t,b) and get returned a data cube (x,y,t,b) with the predicted dates (as passed into predict_labels) on the temporal dimension instead of the original dates. Could you give some more use-case examples? |
Some time ago I've shared with you the python code of the use case implementation, which is here https://github.com/SARScripts/SAR2Cube_use_cases/blob/main/SAR2Cube_Forest_Change.ipynb You can see that we need to fit the curve function over a certain period (2 years of data in this case) but then we need to predict values of a different period (the "testing" one) as well. |
Yes, but the code is mostly undocumented and has a lot of other things in it. I can't exactly figure out what the code does and what part of the code is needed for the openEO use case, sorry.
Okay, so if I understand correctly the issue is that the "parameters" parameter is not a datacube, but an array, right? So the node "array_element (parameters)" is meant to be passed into the "parameters" parameter of fit_curve, right? |
|
Or do you want to predict values without an actual curve_fitting just based on the parameters that are returned for the first fit_curve? Then we'd need two processes: fit + predict. The first fit_curve would be replaced with fit + predict and the second fit_curve would only use predict. |
This! It is indeed what we would need |
|
Okay, thanks for clarifying, that makes things likely a bit more complicated, but I'll come up with another proposal (for two processes!) then. |
|
I've committed a new proposal that splits the fit_curve process into two parts: fit_curve and predict_curve (names TBD). I have to polish the descriptions a lot, but for now it's meant to discuss whether the proposal is feasible and makes sense for many use cases. Here's an example: First of all, the model function should be stored separately, here as an example function stored as a process with the name {
"id": "example_fitting_function_eurac",
"parameters": [
{
"name": "x",
"schema": {
"type": "number"
}
},
{
"name": "parameters",
"schema": {
"type": "array",
"items": {
"type": "number"
}
}
}
],
"process_graph": {
"765a9m5jr": {
"process_id": "pi",
"arguments": {}
},
"a0": {
"process_id": "array_element",
"arguments": {
"data": [
"parameters"
],
"index": 0
}
},
"a1": {
"process_id": "array_element",
"arguments": {
"data": {
"from_parameter": "parameters"
},
"index": 1
}
},
"a2": {
"process_id": "array_element",
"arguments": {
"data": {
"from_parameter": "parameters"
},
"index": 2
}
},
"dzcmqylf5": {
"process_id": "multiply",
"arguments": {
"x": 2,
"y": {
"from_node": "765a9m5jr"
}
}
},
"viadlidxt": {
"process_id": "multiply",
"arguments": {
"x": 2,
"y": {
"from_node": "765a9m5jr"
}
}
},
"l3wirzu5v": {
"process_id": "divide",
"arguments": {
"x": {
"from_node": "dzcmqylf5"
},
"y": 362.25
}
},
"az83u1fgl": {
"process_id": "divide",
"arguments": {
"x": {
"from_node": "viadlidxt"
},
"y": 362.25
}
},
"4kceskraz": {
"process_id": "multiply",
"arguments": {
"x": {
"from_node": "l3wirzu5v"
},
"y": {
"from_parameter": "x"
}
}
},
"4kt3ezdz5": {
"process_id": "multiply",
"arguments": {
"x": {
"from_node": "az83u1fgl"
},
"y": {
"from_parameter": "x"
}
}
},
"kydj4bwy3": {
"process_id": "sin",
"arguments": {
"x": {
"from_node": "4kceskraz"
}
}
},
"o7s99j9pd": {
"process_id": "cos",
"arguments": {
"x": {
"from_node": "4kt3ezdz5"
}
}
},
"82rlj4j02": {
"process_id": "multiply",
"arguments": {
"x": {
"from_node": "a2"
},
"y": {
"from_node": "kydj4bwy3"
}
}
},
"e19f54kwn": {
"process_id": "multiply",
"arguments": {
"x": {
"from_node": "a1"
},
"y": {
"from_node": "o7s99j9pd"
}
}
},
"f9xv6wbqn": {
"process_id": "add",
"arguments": {
"x": {
"from_node": "a0"
},
"y": {
"from_node": "e19f54kwn"
}
}
},
"rjb4pmv5a": {
"process_id": "add",
"arguments": {
"x": {
"from_node": "f9xv6wbqn"
},
"y": {
"from_node": "82rlj4j02"
}
},
"result": true
}
}
}Here's an example workflow based on @clausmichele's image above (if I understood it correctly as the actual values/inputs are missing partially from the image): {
"process_graph": {
"1": {
"process_id": "load_collection",
"arguments": {
"id": "COPERNICUS/S2",
"spatial_extent": null,
"temporal_extent": [
"2015-01-01T00:00:00Z",
"2018-01-01T00:00:00Z"
],
"bands": null
}
},
"2": {
"process_id": "filter_temporal",
"arguments": {
"data": {
"from_node": "1"
},
"extent": [
"2015-01-01T00:00:00Z",
"2017-01-01T00:00:00Z"
]
}
},
"3": {
"process_id": "filter_temporal",
"arguments": {
"data": {
"from_node": "1"
},
"extent": [
"2017-01-01T00:00:00Z",
"2018-01-01T00:00:00Z"
]
}
},
"4": {
"process_id": "fit_curve",
"arguments": {
"data": {
"from_node": "2"
},
"function": {
"process_graph": {
"1": {
"process_id": "example_fitting_function_eurac",
"arguments": {
"x": {
"from_parameter": "x"
},
"parameters": {
"from_parameter": "parameters"
}
},
"result": true
}
}
},
"parameters": [
1,
1,
1
],
"dimension": "t"
}
},
"5": {
"process_id": "predict_curve",
"arguments": {
"data": {
"from_node": "3"
},
"parameters": {
"from_node": "4"
},
"function": {
"process_graph": {
"1": {
"process_id": "example_fitting_function_eurac",
"arguments": {
"x": {
"from_parameter": "x"
},
"parameters": {
"from_parameter": "parameters"
}
},
"result": true
}
}
},
"dimension": "t"
},
"result": true
}
}
}The example likely misses the "labels" in "predict_curve" as they are missing from the example from @clausmichele. It's not clear for which values predictions are computed. |
|
This seems to work, so shall we merge, @clausmichele ? |
|
Yes, for me it's fine! |


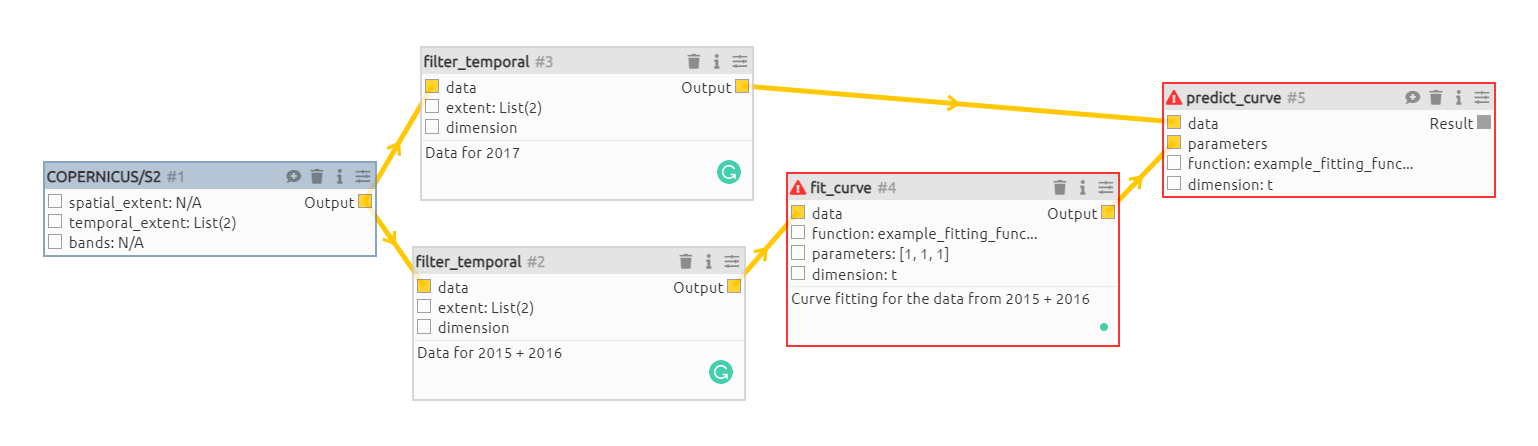
A first draft for the curve fitting process. Name
fit_curveand highly inspired by scipy.curve_fit. This is mostly to spur discussions rather than a final proposal.Related to #231