#1301 changed BASIL input parameters#1360
Conversation
fc5e9b7 to
5407e8c
Compare
Modules/SubModule_Population/xASL_wrp_CreatePopulationTemplates.m
Outdated
Show resolved
Hide resolved
 jan-petr
left a comment
jan-petr
left a comment
There was a problem hiding this comment.
Answering most comments. But I still need to check units for ABV and the correct quantificaiton.
Modules/SubModule_Population/xASL_wrp_CreatePopulationTemplates.m
Outdated
Show resolved
Hide resolved
|
Okay, so between step 3 and step4 (where we do spatial regularization) the maps become weird, the FTISS output from FSL (used later on to obtain CBF.nii by dividing it by M0) becomes weird: But this doesn't mean there's some kind of error, I think that all is working very well, maybe we just need to turn off the bSpai |
|
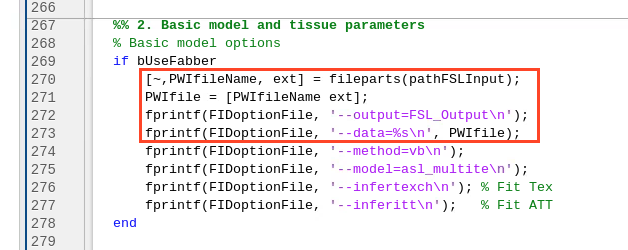
Okay, for fabber quantification, it was giving this error: So, comparing to an older branch (#927), I realised that the fabber command line should be as simple as 'fabber_asl -@ txt_path ' and then all of the options should be inside the txt, including the output path and the PWI input path as well. I've now added a new input to the xASL_sub_FSLOptions: ... so that we can have these two commands inside the txt file: Additionally, since we don't need nothing else than this orange command line, we can define these cmdlineOptions (in green) only for non-Fabber quantification: |
2f202a3 to
be1deb8
Compare






Linked issue
Close #1301